Latest News from the NCDIR
Research Spotlight – Nanobodies targeting SARS-CoV-2 resistant to VOC
A new article published in eLife of work by NCDIR scientists has generated a huge repertoire of high-affinity nanobodies targeting all major regions of SARS-CoV-2 Spike protein.
The work carried out by members from the Rout, Chait and Aitchison Labs have managed to identify from their repertoire of nanobodies, highly synergistic combinations of nanobodies that increase the potency of neutralization, evade escape mutations leading to SARS-CoV-2 variants of concern (VOC) by employing novel mechanisms of action on the SARS-CoV-2 Spike protein.
The large repertoire of high affinity and highly specific nanobodies serves also to make available classes of nanobodies, whose binding covers a significant area of the Spike protein, resulting in nanobdoies that are still *active* against VOC with the potential to also be effective against emerging VOCs.
To read more about this groundbreaking work, read the full article here.
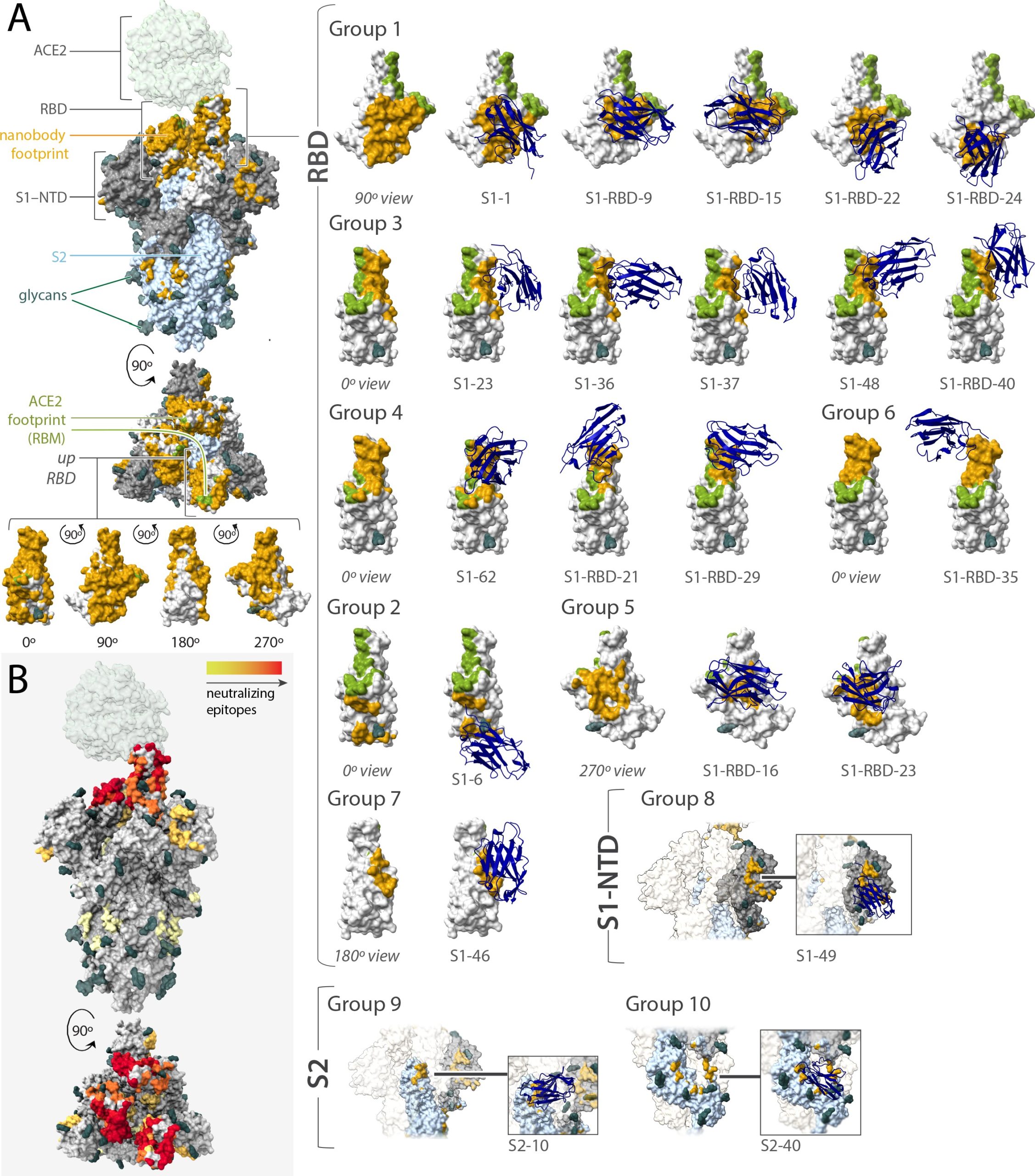
(A) Mapping the binding footprint of the Nanobody classes generated in this study against SARS-CoV-2 reveals significant binding coverage of nanobodies on Spike. (B) the neutralization strength of epitope regions on SARS-CoV-2 spike bound by nanobodies generated in this study, where red is strong neutralization and yellow is weak.
Research Spotlight – Structural basis for transcription complex disruption by the Mfd translocase
A landmark structural biology paper on understanding how a transcription-coupled repair protein disassembles a stalled transcription complex was recently published from the lab of Seth Darst and Elizabeth Campbell with the help of NCDIR scientist Dom Olinares from Brian Chait’s Lab.
Transcription is an essential and multi-step process wherein genetic information encoded in DNA is transcribed into RNA by the RNA polymerase (RNAP) complex. When the bacterial transcription elongation complex (EC) gets stuck on damaged DNA, an ATP-powered, transcription repair protein known as Mfd binds to the EC and displaces the stalled RNAP complex from the DNA.
To capture the structural details involved in the activity of Mfd, the group used single-particle cryo-EM. However, they initially encountered issues with sample preparation—the starting conditions used did not yield any Mfd-bound transcription complexes.
“We performed a sample quality check using the native mass spectrometry (MS) platform that was developed here at NCDIR and we mainly did not observe Mfd bound to the EC. So either Mfd was not binding at all or it quickly falls off after binding to RNAP. These are not optimal for cryo-EM analysis. To efficiently determine conditions that yield intact Mfd-bound complexes, we screened for samples using the same native MS platform by varying incubation times, DNA/RNA sequences, and Mfd preparation. It turns out that all these parameters were critical,” according to Dom.
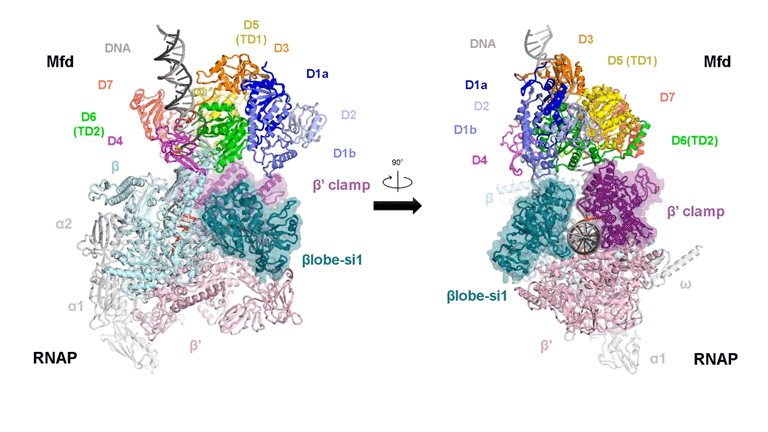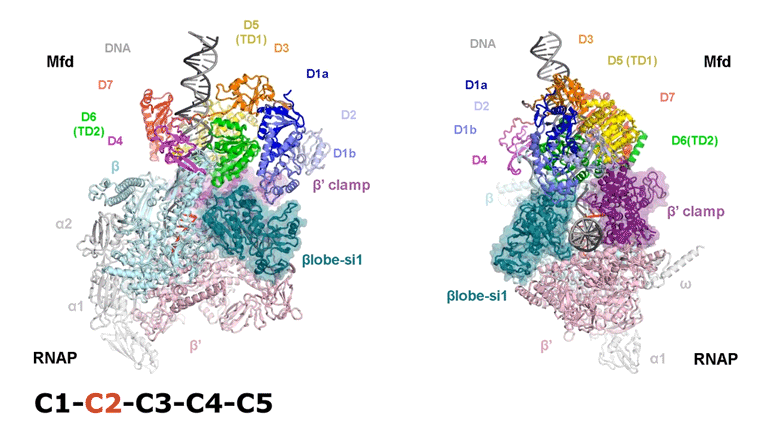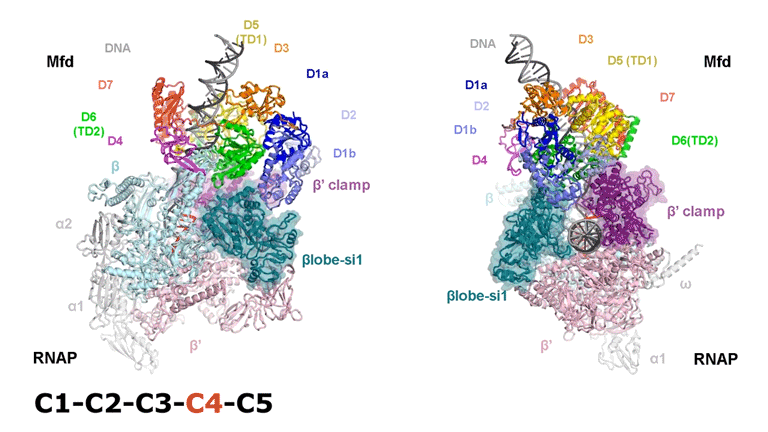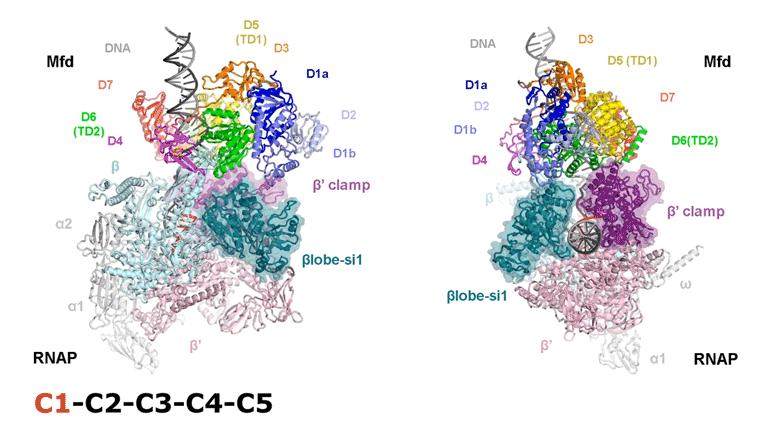
The video shows two orthogonal views of the Mfd-bound complex and cycles through the five structures of the Mfd-nucleotide hydrolysis (C1 → C2 → C3 → C4 → C5). At C2 → C3, Mfd opens the RNAP clamp and at C4 → C5, Mfd twists the βlobe-Si1 domain. The multiple domains of Mfd are colored and indicated accordingly (Kang et. al., 2021).
The native MS-based screening generated the best combination of sample conditions for preparing intact and homogeneous Mfd-bound transcription complexes for cryo-EM analysis. The result? Seven distinct structures that capture how Mfd engages and disassembles the transcription complex at different ADP/ATP states. Overall, the structural intermediates delineate how the initial interaction of Mfd with the stalled complex triggers a stepwise series of dynamic conformational changes, culminating in the stable engagement of Mfd with the EC and then ATP-powered disassembly of the stalled complex (see animation).
The paper is freely accessible here:
https://elifesciences.org/articles/62117
Kang JY, Llewellyn E, Chen J, Olinares PDB, Brewer J, Chait BT, Campbell EA, Darst SA. “Structural basis for transcription complex disruption by the Mfd translocase.” Elife. 2021 Jan 22;10:e62117. PMID: 33480355.
Research Spotlight—Mass spectrometry-based diagnostic and screening platform for optimizing sample preparation in single particle cryo-EM
A native mass spectrometry (nMS)-based platform developed at the NCDIR for evaluating and optimizing sample quality for single particle cryogenic microscopy (cryo-EM) is featured in the current issue of the journal Structure (https://www.cell.com/structure/home).
The exponential increase in high-resolution biomolecular structures determined by cryo-EM is yielding unprecedented structural insights into cellular processes. Despite its extraordinary capabilities, one major bottleneck in single-particle cryo-EM involves sample preparation and the need to have an efficient means for assessing the quality of samples prior to committing the substantial resources, time and effort needed for a typical cryo-EM analysis. To address this need, Dom Olinares, a Research Associate in Brian Chait’s lab at the NCDIR worked with structural biologists to develop an nMS platform that readily integrates into the cryo-EM sample preparation workflow providing rapid feedback on sample integrity/purity and optimal preparation conditions.
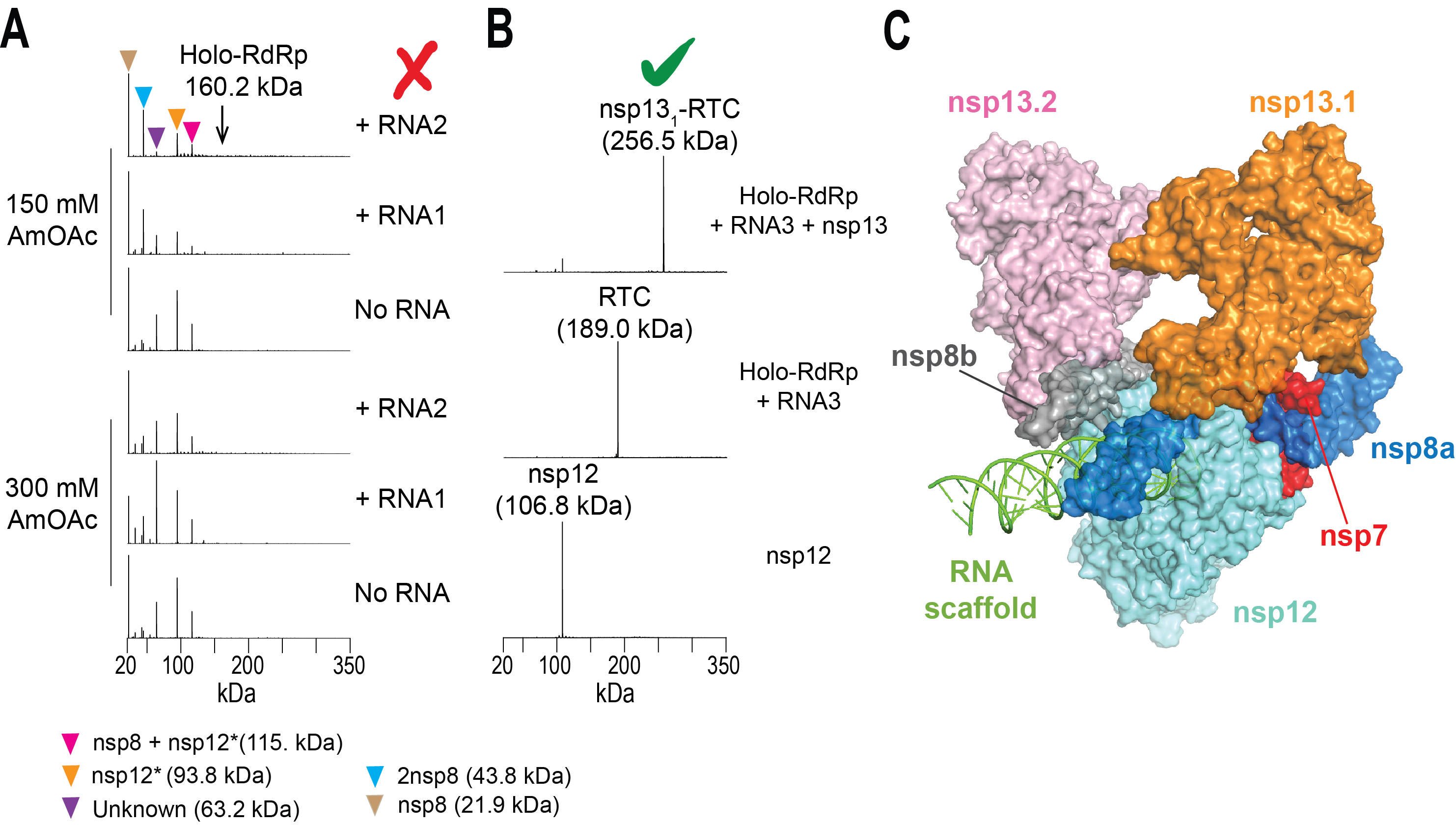
Figure. nMS-based screening to assemble the intact SARS-CoV-2 helicase-RTC. (A) Screening initial conditions for RTC with varying RNA scaffolds and ammonium acetate (AmOAc) concentrations did not form any assembled complex. Detection of unassembled subunits and a truncated nsp12 indicated issues with nsp12 expression. (B) nMS results after optimizing expression of the full-length nsp12 and subsequent reconstitution of RTC. Incubation of the assembled RTC with the nsp13 helicase yielded a single peak corresponding to nsp13-RTC at 1:1 stoichiometry. (C) cryo-EM structure of the nsp13-bound RTC (PDB ID: 6xez) (Olinares et. al., 2020).
Because nMS enables accurate mass analysis of intact protein complexes, it is well-suited for routine evaluation of the composition, integrity, and homogeneity of samples prior to their plunge-freezing on EM grids. In addition, once the EM data is collected, the nMS spectra acquired from the screening provide critical information on subunit composition, stoichiometry, protein modifications, and bound cofactors/ligands that serve as key inputs for EM structure reconstruction and validation.
The utility of the nMS-based diagnostic and screening platform was demonstrated through examples that characterized several bacterial transcription complexes as well as the replication-transcription complex (RTC) coupled to nsp13 helicase from SARS-CoV-2. Most of these structural projects started out with reconstitution conditions that did not yield the desired target assemblies for cryo-EM. In each case, with the aid of iterative nMS-based screening, it proved possible to rapidly establish optimized sample conditions that yielded cryo-EM structures at high resolution.
Publication:
Olinares PDB, Kang JY, Llewellyn E, Chiu C, Chen J, Malone B, Saecker RM, Campbell EA, Darst SA, Chait BT. “Native Mass Spectrometry-Based Screening for Optimal Sample Preparation in Single-Particle Cryo-EM.” Structure. 2020 Nov 17:S0969-2126(20)30413-5. PMID: 33217329.
The full text of the paper is freely available here:
https://www.cell.com/structure/fulltext/S0969-2126(20)30413-5
